However, the efficiency of mapping algorithms has improved substantially over the past decade, while software for QC has received far less attention. In comparison, the time and computation required for QC should be negligible. With the exception of sequence assembly applications, read mapping should be the most computationally expensive step early in analysis pipelines. Common QC tests include counting relative frequency of nucleotides in each position of a set of reads to detect potential deviations from expected frequencies, summarizing the distribution of Phred 10 quality scores to identify base positions with globally low quality (suggesting degeneration in the sequencing process), and measuring the frequency of sequencing adapters and contaminants that are not expected to be biological DNA from the sample.ĭata that passes specific QC tests then undergoes downstream analysis steps, which may include adapter trimming, filtering contaminants and low-quality reads, and mapping the resulting reads to a reference genome or transcriptome. 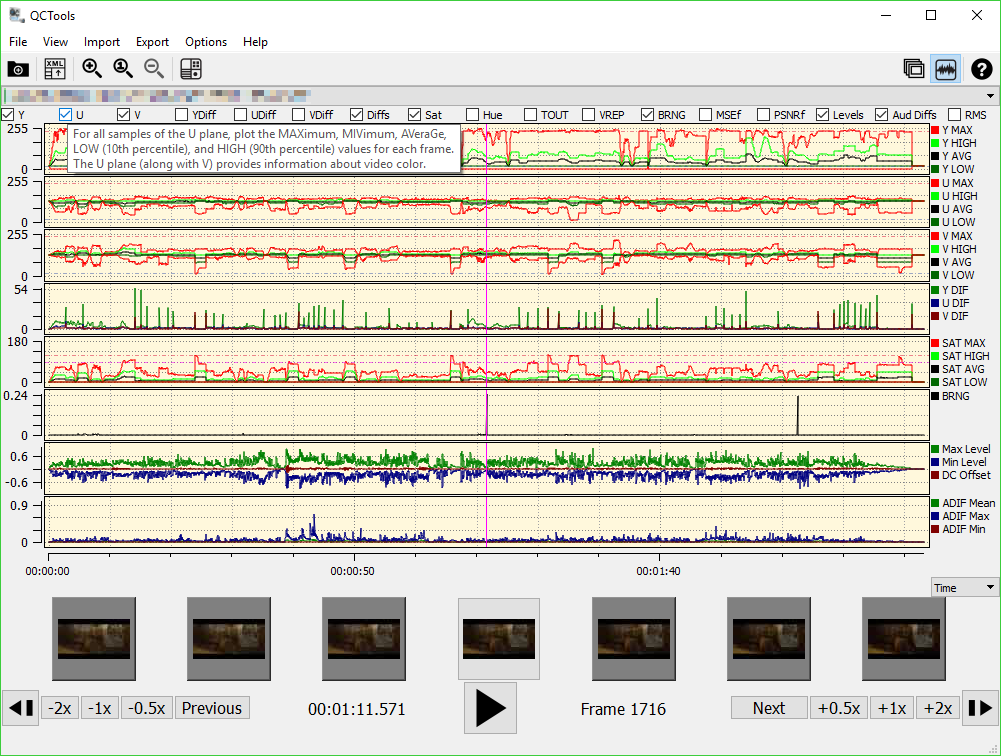 
The QC step measures a set of statistics in a file of sequenced reads to assess if its content matches the experiment expectations and if the data is suitable for downstream analysis. Quality control (QC) is often the first step in high-throughput sequencing data analysis pipelines. New sequencing protocols are constantly being introduced 7, 8, and as the cost of sequencing per base decreases, sequencing data is growing in abundance, dataset size, and read length 9. High-throughput sequencing is routinely used to profile copy number variations in cancers 1, assemble genomes of microbial organisms 2, 3, quantify gene expression 4, identify cell populations from single-cell transcriptomes in a variety of tissues 5, and track epigenetic changes in developing organisms and diseases 6, among numerous other applications. See the authors' detailed response to the review by Weihong Qi See the authors' detailed response to the review by R. (5) The subsection "Falco scales for larger nanopore reads" has been removed, and instead replaced with an additional paragraph on section "Falco is faster than popular QC tools", where the memory usage of each tool in each tested sample is discussed (6) Instructions to report bugs and errors is reported in the "Software availability" section (7) Formatting corrections were performed across the manuscript: "Falco" is now written in uppercase, superfulous line breaks were removed, reference formatting and the usage of the Oxford comma were standardized, links were separated from punctiation and two references were added. (4) The "Methods" section now contains a "system requirements" subsection that describes the memory and disk requirements to run Falco. (3) The "Implementation choices" subsection under "Methods" now highlights that Falco does not contain a user interface, and that Falco was designed for UNIX systems. (2) The "Introduction" section includes more detail about quality control applications. The main changes to the manuscript are listed below: (1) The abstract was changed to highlight the memory comparison between QC software tools, and no longer mentions that FastQC does not run on long-read samples. The code changes that relate to the core computations were not altered since version 0.1.0 (used in the previous version of the manuscript), and we have verified that the times reported in Table 3 remain the same in both versions. Falco version 0.2.4 was used ion this revised manuscript. The accompanying code for Falco has also undergone updates for this review. No other table or figured was altered from the first version of the manuscript. Tables 3 and 4 were expanded to include time measurements for FastQC on long-read samples. Changes to the text were made in all sections. This article has been updated to address reviewer responses. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
